 |
pyclustering
0.10.1
pyclustring is a Python, C++ data mining library.
|
 |
pyclustering
0.10.1
pyclustring is a Python, C++ data mining library.
|
Class represents Elbow method that is used to find out appropriate amount of clusters in a dataset. More...
Public Member Functions | |
| def | __init__ (self, data, kmin, kmax, **kwargs) |
| Construct Elbow method. More... | |
| def | process (self) |
| Performs analysis to find out appropriate amount of clusters. More... | |
| def | get_amount (self) |
| Returns appropriate amount of clusters. | |
| def | get_wce (self) |
Returns list of total within cluster errors for each K-value, for example, in case of kstep = 1: (kmin, kmin + 1, ..., kmax). | |
Class represents Elbow method that is used to find out appropriate amount of clusters in a dataset.
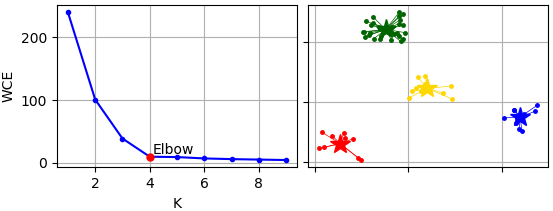
The elbow is a heuristic method of interpretation and validation of consistency within cluster analysis designed to help find the appropriate number of clusters in a dataset.Elbow method performs clustering using K-Means algorithm for each K and estimate clustering results using sum of square erros. By default K-Means++ algorithm is used to calculate initial centers that are used by K-Means algorithm.
The Elbow is determined by max distance from each point (x, y) to segment from kmin-point (x0, y0) to kmax-point (x1, y1), where 'x' is K (amount of clusters), and 'y' is within-cluster error. Following expression is used to calculate Elbow length:
\[Elbow_{k} = \frac{\left ( y_{0} - y_{1} \right )x_{k} + \left ( x_{1} - x_{0} \right )y_{k} + \left ( x_{0}y_{1} - x_{1}y_{0} \right )}{\sqrt{\left ( x_{1} - x_{0} \right )^{2} + \left ( y_{1} - y_{0} \right )^{2}}}\]
Usage example of Elbow method for cluster analysis:
By default Elbow uses K-Means++ initializer to calculate initial centers for K-Means algorithm, it can be changed using argument 'initializer':
| def pyclustering.cluster.elbow.elbow.__init__ | ( | self, | |
| data, | |||
| kmin, | |||
| kmax, | |||
| ** | kwargs | ||
| ) |
Construct Elbow method.
| [in] | data | (array_like): Input data that is presented as array of points (objects), each point should be represented by array_like data structure. |
| [in] | kmin | (int): Minimum amount of clusters that should be considered. |
| [in] | kmax | (int): Maximum amount of clusters that should be considered. |
| [in] | **kwargs | Arbitrary keyword arguments (available arguments: ccore, initializer, random_state, kstep). |
Keyword Args:
True then C++ implementation of pyclustering library is used (by default True).None, current system time is used).1). | def pyclustering.cluster.elbow.elbow.process | ( | self | ) |